Abstract
We present a method for pharmacokinetic modeling of distributions of 111In-labeled monoclonal antibodies (mAbs) on individual pixels of planar scintillation-camera images. Methods: The method is applied to 2 sets of clinical whole-body images, each consisting of 6 consecutive images acquired over a week. Quantification is performed on a pixel basis, yielding images in units of Bq/pixel. The images acquired on the different occasions are registered using a nonrigid method, and for each pixel location a time–activity curve is obtained for which kinetic modeling is performed. The 111In-mAb is assumed to be located in either the vascular or the extravascular space. The vascular content is assumed to follow the global blood kinetics as determined from blood samples, together with a model parameter α that describes the fraction of the whole-body blood volume present in the particular pixel. The rate of change of the extravascular compartment is described by a linear 1-tissue-compartment model with 2 rate constants, K′1 and k2, reflecting extravasation and washout, respectively. The model is optimized for each pixel position with regard to the values of the 3 parameters (α, K′1, and k2), resulting in 3 parametric images. From these, images of the cumulated activity in vascular and extravascular spaces are calculated, as is an image of the rate-constants ratio, which is closely related to the volume of distribution. Results: The resulting parametric images are analyzed in terms of the appearance of the time–activity curves at various locations. Results also include interpretation of the parametric images in their clinical context, and the location of regions that exhibit high extravasation and a low washout rate is compared with confirmed malignant sites. Conclusion: Parametric imaging allows the study and analysis of the spatial and temporal distributions of mAbs simultaneously. Parametric imaging enhances regions where the pharmacokinetics differ from the surrounding tissue and provides a tool to detect and locate unexpected kinetic behavior, which is sometimes characteristic of malignant tissue. For dosimetry in radionuclide therapy, parametric imaging offers a less biased means of analyzing serial mAb images than traditional region-of-interest–based analysis.
Time series of functional images can be analyzed using various types of pharmacokinetic models to obtain information about the rate of absorption and excretion of pharmaceuticals. The parameters of such models depend on various physiologic processes in tissue and on the biochemical properties of the tracer molecule, such as blood and lymph flow, transcapillary exchange permeability, and binding affinity (1–3). Although these physiologic processes cannot be determined in detail using macroscopic modeling, important information about the tissue characteristics can be obtained and may help to improve our understanding of the underlying physiologic and pathophysiologic mechanisms. In radioimmunotherapy, there are also dosimetric reasons for performing pharmacokinetic modeling (4–6). From the values of the kinetic parameters, quantities such as the residence time or cumulated activity can be calculated. Furthermore, the establishment of a pharmacokinetic model that includes typical ranges of transfer rate constants for organs at risk would be of interest both for optimizing radiopharmaceutical activity, such as regarding the organs at risk and therapeutic efficacy, and for optimizing strategies to manipulate biokinetics, such as extracorporeal affinity adsorption (7,8).
Pharmacokinetic modeling can be applied at different levels of detail, using different kinds of compartments that describe the temporal behavior of the unit of interest. Compartments may consist of the entire body, separate organs, subunits of organs, or other functional units that are expected to exhibit a similar time-course variation (4–6,9). Determining the activity preceding kinetic analyses often relies on image-based activity quantification. Regions of interest (ROIs) for the compartments are usually defined manually, and the sum or average within the ROIs is used for the subsequent pharmacokinetic analysis.
Following the examples of brain studies using PET and SPECT, pharmacokinetic analysis can be applied at a pixel or voxel level (10–12). Provided that the images from each scan are spatially registered, the variation in time can be studied for each pixel position. Kinetic parameter values are determined for each pixel, resulting in parametric images. When one uses ROI-based time–activity data, information about any unexpected kinetic heterogeneities within the ROIs may not be revealed, such as if a subvolume within an organ exhibits different kinetics. Parametric images may thus provide less biased information about the pharmacokinetic distribution, because no assumption is made about the location of the functional units before the kinetic analysis. Rather, the parametric images can be used to outline compartments that exhibit a uniform kinetic pattern. Furthermore, parametric images provide pharmacokinetic information that can be analyzed visually, facilitating a visual understanding of the pharmacokinetic and providing a tool for diagnostic applications to detect and locate unexpected kinetic behavior, which is sometimes characteristic of malignant tissue.
For monoclonal antibody (mAb) images, a pixel-based kinetic analysis was developed by Granowska et al. to improve radioimmunodetection (13). A significant positive change in count density between 2 planar images was calculated on a pixel basis and analyzed using statistical methods. The basic assumption was that tumor mAb uptake increases with time, whereas the blood pool, nonspecific uptake, and other tissue activity decrease. Kolbert et al. (14) developed a method of generating parametric rate images from several planar mAb images acquired over an hour, by fitting a linear expression to the count data and mapping the value of the slope. They compared rate images from several injections and also made interpatient comparisons.
The aim of this work was to investigate the feasibility of pixel-based pharmacokinetic modeling for serial mAb images by incorporating data on the time course of blood activity into a pixel-based compartmental model. A further aim was to clinically validate the parametric images qualitatively. To our knowledge, pixel-based pharmacokinetic modeling that includes a description of the blood supply is new in the modeling of mAb distributions. This application relies on our previous work developing techniques for image-based activity quantification and registration (15,16).
MATERIALS AND METHODS
Data Acquisition
Patient data were obtained from a clinical trial using 111In-labeled, biotinylated rituximab (Mabthera; Roche), a chimeric monoclonal B-cell–specific anti-CD20 antibody. Details have been published elsewhere (8). Data from 2 patients were used in this study. Patient 1 had a transformed mucosa-associated lymphoid tissue lymphoma, stage III, originating in the stomach, with tumor manifestations in the mediastinum, abdomen, and retroperitoneal nodes, which were enlarged. Patient 2 had a transformed follicular lymphoma, stage III, with enlarged cervical, retrocrural, paraaortic, iliac, and inguinal nodes and with the hepatoduodenal ligament affected. The activity administered to patients 1 and 2 was 156 and 153 MBq, respectively, measured in a calibrated well chamber (CRC-15; Capintec). The labeling efficiency was determined by instant thin-layer chromatography and high-performance liquid chromatography to be 93% and 97% for patients 1 and 2, respectively. The labeling procedure also included a step in which diethylenetriaminepentaacetic acid (DTPA) was added to bind any free 111In. Blood samples were collected at 5 and 45 min and at 3, 6, 24, 48, 72, 144, and 168 h after injection. The samples were weighed, and the activity was measured in a calibrated well-type NaI(Tl) detector (1282 CompuGamma CS; Wallac). Anteroposterior whole-body images were acquired on 6 occasions: 1, 24, 48, 72, 144, and 168 h after injection (Fig. 1). A dual-head SPECT system equipped with an x-ray unit for simultaneous anatomic imaging (Discovery VG; GE Healthcare) was used. Imaging was performed in a 15% energy window centered over the 111In 245-keV photopeak, in a 256 × 1,024 matrix with a pixel width of 2.2 mm. A transmission image used for pixel-based attenuation correction was also acquired using the x-ray unit in scanogram mode. Based on the geometric mean of anteroposterior images, activity images were calculated in units of Bq/pixel (16). Images were registered at several steps for quantifying activity and for coaligning the time series of the images. The registration method was designed for whole-body images and included nonrigid transformations for each leg, the trunk, the shoulders, and the head, with mutual information as a similarity measure (15).
Image data for patient 1. (A) Attenuation map obtained from CT scanogram image. (B) Geometric mean of anteroposterior raw-count images, showing 3 of total of 6 images, acquired at 1.5, 48, and 163 h after injection. (C) Quantitative images in units of MBq/pixel spatially registered to attenuation map.
Kinetic Modeling
A particular difficulty with planar images, compared with 3-dimensional volumetric data from PET or SPECT, is that the depth dimension is unresolved. The measurement provides no information about the volume corresponding to the measured activity, and volume varies with pixel position. This difficulty complicates the use of a blood concentration curve as an input function, because the blood and image data cannot be directly normalized to a common volume. The relation between the activity concentrations that drive the transcapillary exchange and the measured pixel values is therefore described in some detail below.
In the quantified images, each pixel value is assumed to represent the activity contained in a volume element oriented along the projection line, with the cross-section given by the pixel dimensions ΔxΔy (cm2) and height determined by the total thickness, Ti (cm), of the tissue containing activity at location i (with coordinates (xi,yi)). The total activity, ai(t) (Bq), corresponding to pixel i at different times, t, can be described as the sum of the activity contained in 1 vascular and 1 extravascular compartment, according to: Eq. 1where av,i(t) and aev,i(t) are the activities (Bq), cv(t) and cev,i(t) are the activity concentrations (Bq/mL), and vv,i and vev,i are the volumes (mL) of the vascular and extravascular compartments, respectively. The total volume of the tissue containing activity is vi = vv,i + vev,i = ΔxΔyTi (mL), and Vb,i is the fraction of the volume occupied by the vascular compartment, according to:
Eq. 1where av,i(t) and aev,i(t) are the activities (Bq), cv(t) and cev,i(t) are the activity concentrations (Bq/mL), and vv,i and vev,i are the volumes (mL) of the vascular and extravascular compartments, respectively. The total volume of the tissue containing activity is vi = vv,i + vev,i = ΔxΔyTi (mL), and Vb,i is the fraction of the volume occupied by the vascular compartment, according to: Eq. 2
Eq. 2
The vascular activity concentration, cv(t), contains both 111In-mAb and a radiochemical impurity in the form of 111In-DTPA. For the 111In-mAb, it is assumed that the extravasation (i.e., the transfer from blood vessels to tissue) of antibodies is slow in comparison to the circulation, so the concentration, 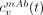 , is equal in all vascular pools. The 111In-DTPA is not modeled explicitly because on the timescale of the mAbs its turnover is fast and its presence is detected only at the early measurement times. Instead, the concentration,
, is equal in all vascular pools. The 111In-DTPA is not modeled explicitly because on the timescale of the mAbs its turnover is fast and its presence is detected only at the early measurement times. Instead, the concentration,  , is assumed to follow the 111In-DTPA kinetics for the whole body, as described in the literature (17). Thus, the activity av,i(t) is determined as:
, is assumed to follow the 111In-DTPA kinetics for the whole body, as described in the literature (17). Thus, the activity av,i(t) is determined as: Eq. 3with
Eq. 3with Eq. 4
Eq. 4 Eq. 5
Eq. 5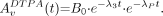 Eq. 6
Eq. 6
The parameter 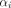 (−) is the fraction of the total-body blood volume, vTB (mL), present in pixel i. Its value is determined in the optimization procedure, and the sum of
(−) is the fraction of the total-body blood volume, vTB (mL), present in pixel i. Its value is determined in the optimization procedure, and the sum of  over the whole image should be near unity for a successful optimization. The functions
over the whole image should be near unity for a successful optimization. The functions  and
and 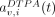 describe the 111In-mAb and 111In-DTPA activities (Bq) contained in vv,i, whereas
describe the 111In-mAb and 111In-DTPA activities (Bq) contained in vv,i, whereas  and
and  are the activities contained in vTB. For
are the activities contained in vTB. For  , biexponential blood kinetics are assumed, with w as the fraction of activity associated with the first component; λ1 and λ2 are the biologic disappearance constants (h−1); and λP is the radioactive decay constant of 111In (h−1). Values of the parameters w, λ1, and λ2 are determined by curve fitting of Equation 5 to the measured activity concentration of blood samples as a function of time. Equation 6 is approximated (17) with a biologic disappearance constant λ3 of 0.415 h−1. The parameters A0 and B0 are the total-body activity at time zero of 111In-mAb and 111In-DTPA, respectively, which are determined from the values of the activity administered and the radiochemical purity measured before administration.
, biexponential blood kinetics are assumed, with w as the fraction of activity associated with the first component; λ1 and λ2 are the biologic disappearance constants (h−1); and λP is the radioactive decay constant of 111In (h−1). Values of the parameters w, λ1, and λ2 are determined by curve fitting of Equation 5 to the measured activity concentration of blood samples as a function of time. Equation 6 is approximated (17) with a biologic disappearance constant λ3 of 0.415 h−1. The parameters A0 and B0 are the total-body activity at time zero of 111In-mAb and 111In-DTPA, respectively, which are determined from the values of the activity administered and the radiochemical purity measured before administration.
The transport of 111In-mAb is derived starting from the formalism used in PET (10,12,18) for a 1-tissue-compartment model, in which the rate of change of the extravascular activity concentration,  (Bq/mL/h), can be described by the differential equation:
(Bq/mL/h), can be described by the differential equation: Eq. 7
Eq. 7
K1,i is the transfer rate constant from vascular to extravascular space normalized to the “tissue” or extravascular volume (mL of plasma/h/mL of extravascular space), k2,i is the rate constant for the washout from extravascular space back to vascular plasma by direct transcapillary passage or indirectly via the lymphatics (h−1),  is the plasma concentration (Bq/mL), and the last term describes the loss due to radioactive decay. The rate of change of the extravascular activity,
is the plasma concentration (Bq/mL), and the last term describes the loss due to radioactive decay. The rate of change of the extravascular activity,  , can then be described by:
, can then be described by: Eq. 8Because the tissue thickness, Ti, cannot be determined from the planar images, and the value of Vb,i varies with the type of tissue represented by the pixel, the value of vev,i cannot be determined. Another extravasation rate constant, K′1,i, normalized to the plasma volume, is therefore defined:
Eq. 8Because the tissue thickness, Ti, cannot be determined from the planar images, and the value of Vb,i varies with the type of tissue represented by the pixel, the value of vev,i cannot be determined. Another extravasation rate constant, K′1,i, normalized to the plasma volume, is therefore defined: Eq. 9with the unit (mL of plasma/h/mL of plasma = h−1), and vpl,i is the plasma volume in pixel i. Equation 8 is then written as:
Eq. 9with the unit (mL of plasma/h/mL of plasma = h−1), and vpl,i is the plasma volume in pixel i. Equation 8 is then written as: Eq. 10where it is assumed that all vascular activity is in plasma so that
Eq. 10where it is assumed that all vascular activity is in plasma so that 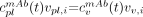 , in agreement with experimental measurements of both the whole blood and the separated plasma activity. Equation 10 is solved analytically for aev,i (Appendix 1). The final expression used for optimization is then obtained by insertion into Equation 1, giving:
, in agreement with experimental measurements of both the whole blood and the separated plasma activity. Equation 10 is solved analytically for aev,i (Appendix 1). The final expression used for optimization is then obtained by insertion into Equation 1, giving: Eq. 11
Eq. 11
A pharmacokinetic quantity of interest is the volume of distribution, Vd, defined as the concentration ratio at equilibrium of extravascular and vascular compartments (10,12). Equilibrium is the condition in which the rate of change of the activity concentration is zero or, when radioactive decay is considered, in which the rate of change corresponds to the radioactive decay. Vd can be interpreted as the volume that the pharmaceutical would need if its concentration were equal to that of the blood concentration and thus reflects the extravascular accumulation of 111In-mAb, which is expected to be related to the amount of binding. For the 1-tissue-compartment model described by Equation 7, Vd is the ratio K1/k2. For the formalism given in Equation 10, the ratio of concentrations cannot be determined. However, by setting the rate of change of the extravascular activity equal to  , the ratio of the extravascular to vascular activity at equilibrium, termed V′d,i, is obtained:
, the ratio of the extravascular to vascular activity at equilibrium, termed V′d,i, is obtained: Eq. 12which is thus Vd,i scaled by the ratio of the extravascular to plasma volumes. It can also be shown that the scaling is inverse to Vb,i and the hematocrit. Thus, although a numeric value of Vd,i cannot be determined, the quantity V′d,i is still of interest for parametric imaging applications.
Eq. 12which is thus Vd,i scaled by the ratio of the extravascular to plasma volumes. It can also be shown that the scaling is inverse to Vb,i and the hematocrit. Thus, although a numeric value of Vd,i cannot be determined, the quantity V′d,i is still of interest for parametric imaging applications.
The cumulated activity, Ã, is a quantity of interest for dosimetry applications, describing the total number of decays occurring in a given region. For the extravascular compartment, Ãev,i is obtained by integration of aev,i (Eq. 5A) from zero to infinity, yielding: Eq. 13in units of (Bq × h). For the vascular compartment, Ãv,i is obtained by integration of Equation 3:
Eq. 13in units of (Bq × h). For the vascular compartment, Ãv,i is obtained by integration of Equation 3: Eq. 14
Eq. 14
Optimization and Implementation
To practically handle the sequential image data, a 3-dimensional image volume is generated with 2 dimensions according the matrix of the scan, with the third dimension given by the number of measured time points. First, the pixels outside the patient, as identified from the attenuation map, are excluded. Then, values of K′1,i, k2,i, and  are determined by fitting Equation 11 to the measured time–activity curve of each pixel i in a least-squares sense. No decay correction is applied before optimization. A gradient-expansion algorithm is normally used for optimization, although the Powell algorithm (19,20) has also been tested and shown to give identical results.
are determined by fitting Equation 11 to the measured time–activity curve of each pixel i in a least-squares sense. No decay correction is applied before optimization. A gradient-expansion algorithm is normally used for optimization, although the Powell algorithm (19,20) has also been tested and shown to give identical results.
Initial values of the parameters to be optimized are estimated from data based on procedures normally used in linearization approaches (21). The initial value of  is set to the fraction of the injected activity in the first time-point image, αi ≈ ai(t ≈ 1 h)/(A0 + B0). The initial value of K′1,i is calculated by integration of Equation 10 up to a certain time point τ, using the assumption that at early time points the impact of aev,i is negligible, so that:
is set to the fraction of the injected activity in the first time-point image, αi ≈ ai(t ≈ 1 h)/(A0 + B0). The initial value of K′1,i is calculated by integration of Equation 10 up to a certain time point τ, using the assumption that at early time points the impact of aev,i is negligible, so that: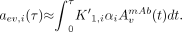 Eq. 15Solving for K′1,i then yields:
Eq. 15Solving for K′1,i then yields: Eq. 16The integral in the denominator is evaluated to:
Eq. 16The integral in the denominator is evaluated to: Eq. 17where τ has been chosen to be the third measured data point, at around 48 h. The initial value of k2,i is calculated by assuming that equilibrium is reached at the last data point, ξ ≈ 170 h, so V′d,i can be approximated by the ratio of the extravascular and vascular activity ratios. Solving Equation 12 for k2,i then yields:
Eq. 17where τ has been chosen to be the third measured data point, at around 48 h. The initial value of k2,i is calculated by assuming that equilibrium is reached at the last data point, ξ ≈ 170 h, so V′d,i can be approximated by the ratio of the extravascular and vascular activity ratios. Solving Equation 12 for k2,i then yields: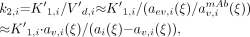 Eq. 18where the DTPA has been disregarded because it has been completely excreted at later time points.
Eq. 18where the DTPA has been disregarded because it has been completely excreted at later time points.
To test the sensitivity to the initial conditions, tests using the same initial values over the whole image have also been made. It was found that results are quite insensitive to the values of k2,i, whereas for K′1,i, initial values of around 0.01–0.1 produce consistent results. Parameter values corresponding to those obtained using the above pixel-based initial estimates were obtained, although there were considerably more pixel positions that did not reach convergence, producing a generally poorer image quality.
To handle noise in the data, a multiresolution filtering strategy has been developed, by which gradually more low-pass filtering is applied at pixel locations where the optimization does not converge. For instance, if a pixel time–activity curve optimization does not converge when no filtering is applied, then the pixel value in each activity image in the time series is replaced by the average of a 3 × 3 pixel neighborhood, and then the optimization is repeated. Average filters are applied with kernels of widths ranging from 1 (i.e., no filtering) up to 19 pixels wide, corresponding to filter widths of 2.2 mm to 4.2 cm. The level of filtering used for successful optimization is stored for each pixel and is used as a measure of the goodness of fit. Furthermore, the squared correlation coefficient between the measured values and the model-predicted values is calculated and mapped for each pixel (19). Thus, 2 images of the goodness of fit are obtained and used as maps of the reliability when interpreting the pixel values of the parametric images.
A computer program that has been developed for analysis of the pharmacokinetics allows the user to interactively navigate a cross-hair in the image and consecutively displays the values of the parameters and of the goodness of fit and also plots the time–activity curves for the selected location. All computer programs were implemented in Interactive Data Language (Research Systems, Inc.) (19).
RESULTS
For the blood curves, the biexponential function (Eq. 5) described the data well, with a correlation coefficient of more than 0.998 for both patients.
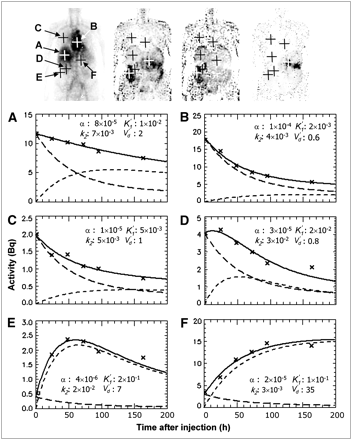
Figure 2 shows parametric images of α, K′1, k2, and V′d for patient 1, resulting from the pharmacokinetic modeling. To illustrate the meaning of the parameters, Figure 3 displays these images together with plots of locations that exhibit representative time–activity curves, subjectively selected by navigating through the image. Figure 3 shows that at time zero all the activity is in the vascular space. The activity value is equal to  , for example 8·10−5·156 MBq ≈ 12 Bq for Figure 4A. At early times, the extravascular activity increases. In terms of the curves, the parameter K′1 influences the slope of the tangent to the initial part of the extravascular time–activity curve, which can be seen mathematically by setting aev to zero in Equation 10. As time passes, the washout from extravascular space gradually has more impact, and at about 50–100 h after injection the slope of the extravascular curve, that is, the rate of change (Eq. 10), decreases toward zero. At later times, the extravascular curve either becomes parallel to the blood curve, corresponding to smaller k2 values, or has a negative slope, corresponding to larger k2 values. The value of V′d is the ratio of the extravascular to vascular activities at equilibrium and can be estimated from the diagrams as the activity ratio at late time points.
, for example 8·10−5·156 MBq ≈ 12 Bq for Figure 4A. At early times, the extravascular activity increases. In terms of the curves, the parameter K′1 influences the slope of the tangent to the initial part of the extravascular time–activity curve, which can be seen mathematically by setting aev to zero in Equation 10. As time passes, the washout from extravascular space gradually has more impact, and at about 50–100 h after injection the slope of the extravascular curve, that is, the rate of change (Eq. 10), decreases toward zero. At later times, the extravascular curve either becomes parallel to the blood curve, corresponding to smaller k2 values, or has a negative slope, corresponding to larger k2 values. The value of V′d is the ratio of the extravascular to vascular activities at equilibrium and can be estimated from the diagrams as the activity ratio at late time points.
Parametric images for patient 1. (A) α: fractional blood volume (−). (B) K′1: normalized extravasation rate constant (h−1). (C) k2: rate constant of washout from extravascular to vascular space (h−1). (D) V′d: extravascular-to-vascular activity ratio at equilibrium, equal to K′1/k2, and closely related to volume of distribution (−). Gray scales are to right of each image, with maximum and minimum values displayed. Patient 1 had tumor in abdomen, corresponding to high-contrast region on V′d image (D).
At top, from left to right, are the parametric images α, K′1, k2, and V′d, showing positions corresponding to time–activity curves for liver (A), heart (B), lung (C), kidney (D), intestinal tract (E), and tumor (F). Decay-corrected activity values from measured images are displayed as crosses. Solid line is fitted total-activity curve (ai(t)), long dashed line is vascular activity (av,i(t)), and short dashed line is extravascular activity (aev,i(t)). Correlation coefficient was between 0.97 and 0.99, and no filtering was used for the positions marked.
Distributions of measures of goodness of fit. (A) Filter width applied for successful optimization of patient 1. Black corresponds to no filtering, and white is maximum level applied (19-pixel-wide average filter). (B) Squared correlation coefficient between measured and fitted time–activity curves of patient 1; black implies high correlation. (C) Frequency histogram of filter width required for successful time–activity curve optimization of both patients.
Figure 3A displays a time–activity curve at a position in the liver. The value of K′1 is 0.01 h−1. At the later time points, the extravascular activity is higher than the vascular activity, with a V′d of 2. Figures 3B and 3C show time–activity curves of the heart and lung. Both time–activity curves are dominated by the vascular compartment, with slow and low extravasation, as reflected by low values of K′1. Figure 3D shows the time–activity curve for the cortex of the right kidney. The value of K′1 is similar to that of the liver, but the washout rate is somewhat higher, yielding a V′d close to 1. Figure 3E is the time–activity curve for a location selected on the basis of the increased values in the K′1 and k2 images. Both the extravasation and the washout rates are high, with a K′1 of 0.2 h−1and a k2 of 0.02 h−1. The location corresponds to the intestinal tract, but we still do not know the actual tissue associated with this fast turnover. Figure 3F is the time–activity curve for a location that corresponds to the abdominal tumor manifestation. It appears that this region has not yet reached equilibrium at 200 h after injection. K′1 is high and the washout rate is low, yielding a V′d of 35 for the chosen pixel location.
Figures 4A and 4B show distributions of the measures of the goodness of fit, that is, the level of filtering required to obtain convergence, and the squared correlation coefficient. Figure 4C shows a frequency histogram of the number of pixel time–activity curves requiring various levels of filtering. The majority of the pixels, about 80%, require no filtering to obtain convergence. About 3%–5% require average filtering using a width of up to 9 pixels, corresponding to the average from a 2 × 2 cm2 neighborhood, and these pixels are primarily near the patient contour and in the distal legs (Fig. 4A). This could be due to differences in the torsion of the legs between acquisitions—differences that are not properly accounted for by image registration. The correlation coefficient (Fig. 4B) is mostly close to 1.0, although smaller values are obtained at the patient contour because of misregistration. Also noticeable are a point in the abdominal tumor region and an area in the intestinal tract on the patient's left side that exhibit rather poor values of 0.4–0.6.
Figure 5 shows 2 measured raw images and parametric images from patient 2. In general, for the normal organs of patient 2 the shapes of the time–activity curves, separated into vascular and extravascular activity components (not shown), are similar to those of patient 1. For both patients, the α image, Figures 2A and 5C, describing the distribution of blood in the body, is similar to the image obtained at the first time point (Figs. 1B and 5A), with high-intensity areas appearing in blood-rich regions such as the heart, liver, and large vessels. In the K′1 images (Figs. 2B and 5D), high values appear along segments of the patient contour and along the edge of the liver and spleen. In regions where the activity gradient is high, any spatial misregistration between the images acquired at different time points will yield a high variability in the measured time–activity data. If the regions near the patient contour are not considered to be of interest, the value of K′1 may be set to zero, where contour pixels are identified using the attenuation map and applying a mathematic erosion operator (22). Also, to decrease the influence of the pixels where the optimization is poor, the possibility of discriminating the pixels where the correlation coefficient is below a user-defined value is included. This procedure has been applied by setting K′1 to zero for all pixels with a squared correlation coefficient below 0.5 in Figure 5. Furthermore, to be able to visualize V′d for pixels where K′1 is large while k2 approaches zero, such as in the abdominal tumor region of patient 1 in Figure 2, the possibility of replacing k2 by a distance-weighted average of its closest neighbors is included, accomplished by applying a morphologic closing operator (22) to the k2 image.
Patient 2. (A and B) Raw-count geometric-mean images acquired at 1.1 and 164 h after injection. (C–F) Parametric images α (C), K′1 (D), k2 (E), and V′d (F), with lower limbs excluded for space reasons. Arrows a–i indicate positions at which parameter values were obtained: a = liver (α = 5 × 10−5, K′1 = 1 × 10−2, V′d = 1.2), b = heart (α = 6 × 10−5, K′1 = 6 × 10−3, V′d = 0.6), c = lung (α = 1 × 10−5, K′1 = 4 × 10−3, V′d = 0.3), d = kidney (α = 2 × 10−5, K′1 = 3 × 10−2, V′d = 1.4), e = intestinal tract (α = 7 × 10−6, K′1 = 8 × 10−2, V′d = 3), f = paraaortic malignancy (α = 8 × 10−6, K′1 = 9 × 10−2, V′d = 11), and g, h, and i = iliac, inguinal, and cervical nodes, respectively (α = 1 × 10−5, K′1 = 5 × 10−2, V′d = 7–10).
Regions that exhibit homogeneous uptake patterns are distinguishable in the K′1 and k2 images of both patients. However, the images give the impression of being noisy, mostly because of the high variability in tissues that do not accumulate 111In-mAb. The variability is less pronounced in the V′d images (Figs. 2D and 5F), where the artifacts due to spatial mismatch almost disappear. Spatial mismatch and image noise due to low count values produce variability in the data points in the time–activity curves, resulting in greater uncertainty in the parameter values obtained. We have noticed that K′1 and k2 correlate positively and that this correlation leads to reduced variability in V′d. For the K′1 and k2 images, the general intensity of the lungs is similar in the 2 patients, although there is a high variability because of image noise. The appearance of the liver is similar for the 2 patients, with values of V′d between 1.2 and 2. Both patient images exhibit streaks of unexpectedly high K′1 and k2 values in the intestinal tract, although these cancel out in the V′d image, where corresponding streaks are not seen. In both patients, the heart and kidneys exhibit V′d values of less than 1 and around 1, respectively. Interestingly, in the V′d image of patient 2 (Fig. 5F) the kidney (d) appears to be separated into 2 distinct areas with different kinetic rate constants, corresponding to the location of the renal pelvis and cortex. In patient 2, 4 of the sites of enlarged nodes clearly exhibit high V′d values, indicated in Figure 5F. The shape of the time–activity curve (not shown) of the paraaortic malignancy (f) resembles the time–activity curve in Figure 3F, with a V′d of 11 for the selected location. The cervical, iliac, and inguinal nodes (i, g, and h, respectively) present flatter time–activity curves with V′d values of 7–10.
Figure 6 shows images of the cumulated activity separated into its vascular and extravascular components. The Ãv images reflect the distribution of blood and appear similar for the 2 patients. The Ãv images are essentially a scaling of the α images, because in Equation 14 that is the only quantity that varies with image position. The Ãev images differ for the 2 patients and appear similar to the V′d images. For patient 1, values are increased in the abdominal tumor region and also at positions corresponding to the mediastinal tumor manifestations, which were not visualized as well on the V′d image. For patient 2, clearly increased values are obtained in positions corresponding to the 4 enlarged nodes indicated on the V′d image.
Cumulated activity images in patients 1 (A–C) and 2 (D and E). (A) Total cumulated activity (only patient 1 shown). (B and D) Vascular component Ãv. (C and E) Extravascular component Ãev. Ãev images exhibits high contrast values in positions corresponding to confirmed malignant regions, for patient 1 (C) in abdomen and 2 locations in mediastinum, and for patient 2 (E) at cervical, paraaortic, iliac, and inguinal nodes.
DISCUSSION
We have developed a method to calculate whole-body images of the distribution of parameter values of a 1-tissue-compartment model, and we have applied this method to data from patient studies with 111In-mAb. We have shown that compartmental modeling on a pixel level is feasible for this type of data. This tool is good for gaining a comprehensive picture of the pharmacokinetic and for analyzing it at a pixel level without defining ROIs beforehand.
Although the method has been thoroughly tested, the quantitative parameter values presented here are not considered to be fully representative of the tissues: The specific locations were chosen only to illustrate the essence of the parametric images. In an ongoing project, histograms of rate constant distributions in selected ROIs are being quantitatively analyzed in detail and compared with SAAM II–based compartmental modeling (University of Washington). The parameter values obtained have generally been in agreement, although more evaluation remains to be done.
The K′1 and k2 images may be difficult to interpret individually (Figs. 2 and 5), partly because of noise and spatial mismatches that cause the optimization to converge to variable parameter values for some regions. On the V′d images, which describe the extravasated amount of 111In-mAb in relation to the vascular amount, the correlation found between K′1 and k2 leads to reduced noise. The Ãev images (Fig. 6) have a smoother appearance and reveal malignant regions in patient 1 that were difficult to distinguish on the V′d image. V′d and Ãev in combination are concluded to be the quantities best suited for parametric imaging for radioimmunodetection.
The images of cumulated activity also provide quantitative information on the extravascular fraction of total cumulated activity. This information may have implications for dosimetry in situations where the therapeutic effect depends on the proximity of the radiopharmaceutical to the target cells, for example, for radionuclides emitting particles with a short range. Furthermore, for dosimetric applications, parametric images can be of use for defining regions with similar kinetic behavior for a subsequent ROI-based analysis. For example, in Figure 5F, the border of the kidney is clearer than on the activity images, and separation of the cortex and pelvis in this image also appears feasible. Moreover, high-intensity regions seen in the intestinal tract in Figures 2B and 5D but not distinguishable on the activity images can be delineated and then be accounted for as an overlapping region for a subsequent ROI-based pharmacokinetic analysis of the kidneys and liver.
The current 1-tissue-compartment model, although appearing adequate for describing the pharmacokinetic in most image positions, is probably not sufficient for all tissues. Two factors in particular may make the underlying kinetics deviate from a 1-tissue-compartment model, requiring caution in interpreting the parameter values quantitatively. First, because data are obtained from planar images, the parameter values reflect the extravasation and washout rates of all tissue represented by the pixel, including activity in under- or overlying tissues. For the current study, the general background is relatively low, however, making the background kinetics less influential. Also, at least a portion of most organs is nonoverlapping and may provide representative quantitative organ rate constants. The extent to which overlap influences the bias and precision of the parameter values is currently being investigated. Furthermore, it will be of interest to implement the modeling in 3-dimensional SPECT or PET images.
Second, the underlying assumption is that extravasation and washout are governed by mechanisms that behave similarly for all tissues and organs and are linear with the supply of 111In-mAb. For most normal tissues and organs, it is mainly passive mechanisms that are responsible for the transport. Tumors are, however, expected to be capable of binding, and in such cases the disappearance from the extravascular compartment is no longer linear with the total amount of 111In-mAb because not all is available for washout. The parameter k2 of the current formulation then underestimates the true washout rate by the lymphatics. Hence, the parameter V′d becomes high and, in accordance with the high observed values, is unlikely to be due to passive transport only. Furthermore, if the binding is saturable because of a limited number of binding sites, the kinetics become nonlinear. A model description that separates the extravascular compartment into a free compartment and a bound compartment (including saturation) would probably be more appropriate for describing the kinetics of the tumors, as is seen in Figure 4B, in which 2 regions, one near the abdominal tumor and another in the intestinal tract, show low correlation values. Likewise, in Figures 3E and 3F one can envisage that a more complex model would have provided a better fit at later time points. However, at this stage, and given the few data points, the current model was chosen to obtain stability.
A special concern in mAb parametric imaging is that turnover is comparatively slow, on the order of days or even weeks. Parametric imaging in general has been applied mostly in situations where the physiologic process of interest is relatively fast, so the sequential imaging can be performed with the patient in position. Calculation of mAb parametric images requires registration of the images from different time points. A potential source of error is then spatial mismatch, which causes inaccurate parameter values, particularly at locations with a pronounced signal gradient. The registration method applied has shown good accuracy (15), and visual inspection of the registrations shows that generally they appear to be accurate. However, organ movement within the body is not accounted for and generates mismatch errors, for example, the rim at the top of liver in Figures 2B and 5D.
Concerning optimization, convergence is reached for about 80% of the pixel positions without filtering (Fig. 4). However, the advantage of gradual filtering is that parameter values are obtained for all pixels, thus producing a better image quality than would be obtained if the remaining 20% were not calculated. Also, the data-based initial estimates produce good and stable solutions, improving image quality.
CONCLUSION
Compartmental modeling on a pixel level is feasible for planar mAb images, provided that accurate image-based activity quantification and image registration are performed. The resulting parametric images, V′d and Ãev in particular, are believed to have a potential for use in radioimmunoimaging and radioimmunotherapy.
APPENDIX
Solution of Differential Equation (Eq. 10)
The rate of change of the extravascular activity,  is described as:
is described as: Eq. 1A
Eq. 1A Eq. 2A
Eq. 2A Eq. 3AIntegrating this expression, inserting Equation 5, and noting that
Eq. 3AIntegrating this expression, inserting Equation 5, and noting that  , Equation 3A is written as:
, Equation 3A is written as: Eq. 4AThe expression for
Eq. 4AThe expression for  is then obtained as:
is then obtained as: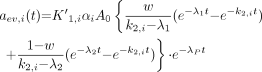 Eq. 5A
Eq. 5A
Acknowledgments
We acknowledge valuable comments from Tomas Ohlsson, PhD, who was responsible for the radiochemical work in the patient studies. This work was financially supported by the Swedish Research Council.
Footnotes
-
COPYRIGHT © 2007 by the Society of Nuclear Medicine, Inc.
References
- Received for publication January 15, 2007.
- Accepted for publication April 20, 2007.